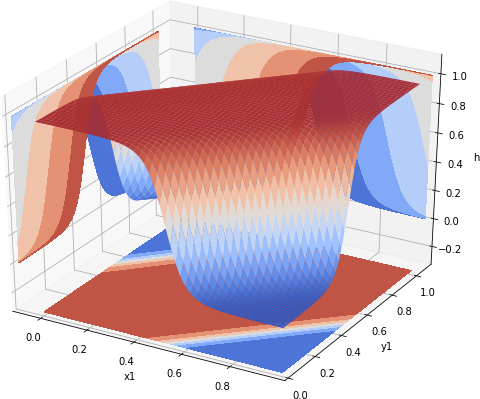
In the last two posts we have shown how to encode, using Tensorflow, a neural network that behaves like the XNOR gate. However, it is a very unusual that we know, ahead of time, weights and biases. A much more common scenario is when we have number of inputs and the corresponding values, and wish to train a neural net to produce for each input the appropriate value. For us the inputs are (0, 0), (0, 1), (1, 0) and (1, 1). The values are 1, 0, 0, and 1. We have seen that a 2 layer deep neural network can emulate XNOR with high fidelity. Thus we could just create a 2 layer deep, 3 neuron network and attempt to train it. To facilitate some level of experimentation we create a function that produces a fully connected neural network layer.
def CreateLayer(X, width):
W = tf.get_variable("W", [X.get_shape()[1].value, width], tf.float32)
b = tf.get_variable("b", [width], tf.float32)
return tf.nn.sigmoid(tf.add(tf.matmul(X, W), b))
The function creates input to a layer of neurons. Using
instead
allows us to specify
inputs (features) as a
matrix. This is often easier than representing inputs as
columns each
high. Also, rather than creating variables directly, with
tf.Variable, we use tf.get_variable. This allows variable sharing, as explained in Sharing Variables. It also can enhance display of the computation graph in TensorBoard, as explained in Hands on TensorBoard presentation.
We also create a training operation and a loss function that allows us to assess how well the current network is doing. Tensorflow offers a whole array of optimizers, but tf.train.AdamOptimizer is often a good choice. In order for the optimizer to push variables to a local minima we must create an optimizer operation. This is done by calling minimize method with a loss function. The loss function tells the optimizer how far it is from the ideal solution. We use the mean of squared errors as our loss function.
def CreateTrainigOp(model, learning_rate, labels):
loss_op = tf.reduce_mean(tf.square(tf.subtract(model, labels)))
train_op = tf.train.AdamOptimizer(learning_rate).minimize(loss_op)
return train_op, loss_op
The above function returns the training and loss operations. The latter is used to track progress of the model towards the optimum. The final piece of code that needs to be written is the training code.
g = tf.Graph()
with g.as_default():
X = tf.placeholder(tf.float32, [None, 2], name="X")
y = tf.placeholder(tf.float32, [None, 1], name="y")
with tf.variable_scope("layer1"):
z0 = CreateLayer(X, 2)
with tf.variable_scope("layer2"):
z1 = CreateLayer(z0, 1)
with tf.variable_scope("xnor"):
training_op, loss_op = CreateTrainigOp(z1, 0.03, y)
init_op = tf.global_variables_initializer()
saver = tf.train.Saver()
X and y (line 3-4) are placeholders, which are going to be seeded with inputs and desired outputs. We specify the first dimension to be None to allow for arbitrary number of rows. In lines 5 – 10 we create a model. It consists of two, fully connected layers. The first layer has 2 neurons, the second consists of a single neuron. X is the input to the first layer, while the output of the first layer, z0, is the input to the second layer. The output of the second layer, z1 is what we wish to train to behave like the XNOR gate. To do so, in lines 9 and 10 we create a training operation and a loss op. Finally we create an operation to initialize all global variables and a session saver.
writer = tf.summary.FileWriter("/tmp/xnor_log", g)
loss_summ = tf.summary.scalar("loss", loss_op)
Before we run the training step we create a summary writer. We are going to use it to track the loss function. It can also be used to track weights, biases, images, and audio inputs. It also is an invaluable tool for visualizing data flow graph. The graph for our specific example is shown in Fig 1.
 Fig 1. Data flow graph as rendered by tensorboard
Fig 1. Data flow graph as rendered by tensorboard
In order to train our model we create two arrays representing features and labels (input values and the desired output). The training itself is done for 5,000 steps by the for loop. We feed the session all inputs and desired values, and run a training operation. What this does it runs the feed forward steps to compute z1 for the given inputs, weights and biases. These are then compared, using the loss function to the ideal responses, represented by y from these Tensorflow computes contributions all weights and biases make to the loss function. It uses the learning rate 0.03 to adjust them to make the loss smaller.
X_train = np.array([[0, 0], [0, 1], [1, 0], [1, 1],])
y_train = np.array([[1], [0], [0], [1]])
sess = tf.Session(graph=g)
sess.run(init_op)
for step in xrange(5000):
feed_dict = {X: X_train, y: y_train}
sess.run(training_op, feed_dict=feed_dict)
if step % 10 == 0:
writer.add_summary(
sess.run(loss_summ, feed_dict=feed_dict), step)
save_path = saver.save(sess, '/tmp/xnor.ckpt')
sess.close()
print "Model trained. Session saved in", save_path
Once the training is complete we save the state of the session, close it and print the location of the single session checkpoint. The loss function, as recorded by the summary file writer, and rendered by TensorBoard, is shown in Fig 2.

Fig 2. Loss function plotted by tensorboard.
At the end of the training it the loss function has the value of 0.000051287. It was still dropping but very slowly. In the next post we show how to restore the session and plot the loss function as well as the output of the trained neural network.
Resources
The Jupyter notebook that implements the above discussed functionality is xnor-train.ipynb in the xnor-train project.